Introduction
SSBD is a platform for sharing and reusing bioimaging data. SSBD:database is an added-value database for bioimages and biological dynamics data. It provides a rich set of open resources for analyzing microscopy images and qunatitative data of biological objects, such as single-molecule, cell, tissue, individual, etc., and software tools for analysis. Microscopy images and quantitative biological data are collected from a variety of species, sources, and methods. These include data obtained from both experiments and computational simulations.
SSBD:database shares 244 projects, SSBD shares totally 326 projects, 43.5 TB (2025-07-01)
Find the dataset from the search box above, or see the dataset list on the Resources.
See Citation Policies(PDF) for citation instructions.
For more information, please refer to the following paper:
Koji Kyoda, Hiroya Itoga, Yuki Yamagata, Emi Fujisawa, Fangfang Wang, Miguel Miranda-Miranda, Haruna Yamamoto, Yasue Nakano, Yukako Tohsato, Shuichi Onami,
SSBD: an ecosystem for enhanced sharing and reuse of bioimaging data,
Nucleic Acids Research , Volume 53, Issue D1, 6 January 2025, Pages D1716-D1723,
doi: 10.1093/nar/gkae860, PMID: 39479781; PMCID: PMC11701685.
Share your data
SSBD:database shares selected, highly reusable bioimaging data and biodynamics data. Our curators will invite to share the data from SSBD:repository and accepted papers. Share youre data on SSBD:repository first, or send e-mail to us.
Samples
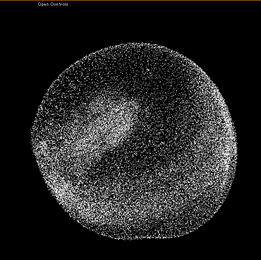
Microscopy Images
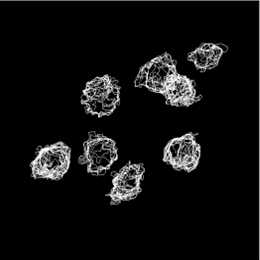
Quantitative Data
News
-
2025-06-02,
New SSBD explainer video released
https://www.youtube.com/watch?v=935KmjA1s9E
Our latest video introduces the key points of the new SSBD paper published in Nucleic Acids Research.
Watch it. -
2024-12-27,
SSBD:database 2024 update
Happy Holidays and Happy New Year!
We are pleased to announce SSBD:database 2024 update.
We released 30 projects in 2024, which include 906 image datasets (5.23TB).
The added projects/datasets have already been announced on https://x.com/ssbd_en .
Again, we thank the contributors for their valuable data.The released projects …
-
2024-12-27,
SSBD:repository 2024 update
Happy Holidays and Happy New Year!
We are pleased to announce SSBD:repository 2024 update.
We released 24 projects (2.5TB) in 2024.
The added projects have already been announced on https://x.com/ssbd_en .
Again, we thank the contributors for their valuable data.The released projects are:
https://ssbd.riken.jp/repository/ssbd-repos-000256
Expression patterns … -
2024-10-31,
New SSBD paper has been published
A new paper on SSBD has been published. It describes the two-tier ecosystem of SSBD:repository and database and the Global Image Data Ecosystem (GIDE), based on a global collaboration.
Koji Kyoda, Hiroya Itoga, Yuki Yamagata, Emi Fujisawa, Fangfang Wang, Miguel Miranda-Miranda, Haruna Yamamoto, Yasue Nakano, Yukako Tohsato, Shuichi Onami, …
-
2024-10-05,
SSBD:repository and SSBD:database integration update
SSBD has revised its data storage policy as follows to ensure proper data collection, metadata annotation, publication, and reuse.
- From SSBD:repository, all original data will be stored and published, and the download and API service will be unified via SSBD:repository
- From SSBD:database, we promote data publication in highly … - Older news...
SNS
Follow @ssbd_en Follow @ssbd_jabioimagram: View in Dataset, View in OMERO